What is VisualPDE?
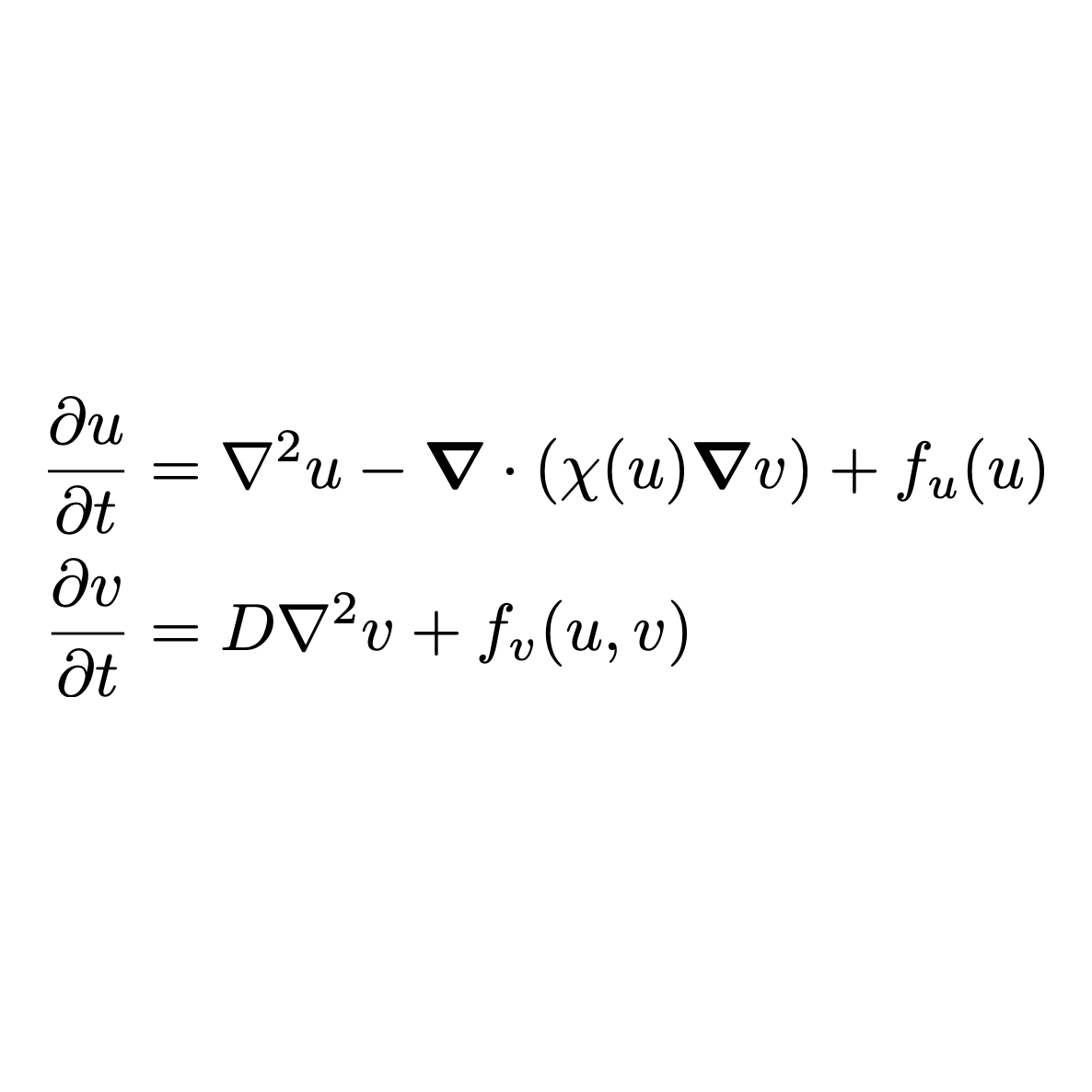
Developing intuition for mathematics can be difficult, whether or not you are a mathematician.
VisualPDE aims to solve this problem.
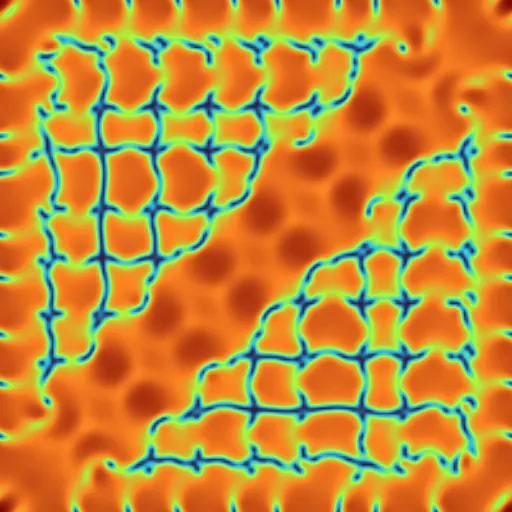

Our lightning-fast simulator makes mathematics visual and interactive on almost any device.
Jump into a simulation to see for yourself.
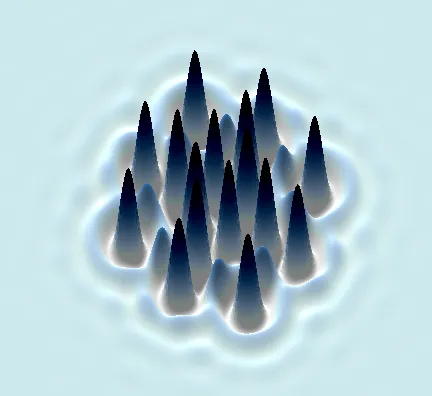



Not a mathematician? No problem.
Try out our interactive Visual Stories for a snapshot of cutting-edge science and mathematics.

Getting started
You can explore the examples, create simulations and share them with a URL, or even copy the code on GitHub to design your own version of VisualPDE. Check out our detailed documentation to dive under the hood.
If you use VisualPDE in your research, we’d be grateful if you could cite our article about the context, design, and applications of VisualPDE in the Bulletin of Mathematical Biology.
Is it free?
Everyone is free to use VisualPDE. For specifics of industrial use, see our licence.
VisualPDE in the wild
VisualPDE is used around the world to teach, engage and interact with mathematics and science. Here are some examples of VisualPDE in action:
- 20+ citing articles on Semantic Scholar
- An interactive logo for the Society for Mathematical Biology
- A simulation-rich blog post by Vatsal Sanjay summarising VisualPDE
- A Plus magazine article and podcast on using VisualPDE to explore research in virus transmission
- A Teaching and Learning Mathematics Online webinar exploring VisualPDE
The team
VisualPDE is a team effort, written and maintained by Benjamin Walker, Adam Townsend and Andrew Krause.




